 These examples were chosen from the PDB Molecule of the Month site. use set cartoon_side_chain_helper, on to only show side chains as sticks.use set surface_quality, 1 to improve the quality of the mesh exported from PyMOL.you can use split_chains to split the protein into chains so you can export each one individually.This means creating the final image with all the realistic lighting, reflections and so on. Apply materials to the model(s) and then render the scene.Once the file is loaded you can set up a scene with some lighting and a camera.If the model is large it might take a minute.You then open Blender and choose file->import X3d Extensible 3D (.wrl).This is a standard file format for representing 3D data. With the model or chains you want visible, choose file->export image as->vrml 2.Once you orient the structure, don’t move it around if you are saving multiple parts so that the models get exported in the same orientation. You can use the split_chains command for this. If the structure is made of multiple chains you might want to split them up so they can be imported as separate objects.Display the structure as you want it to look in the final model i.e.It’s also possible to import pdb files directly into Blender but this seems fairly limited. A more detailed guide will require a video tutorial. This is a brief summary of the method used. 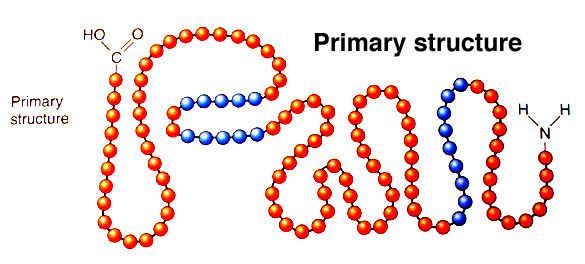
If you’re not interested in the method used skip below to see some examples. 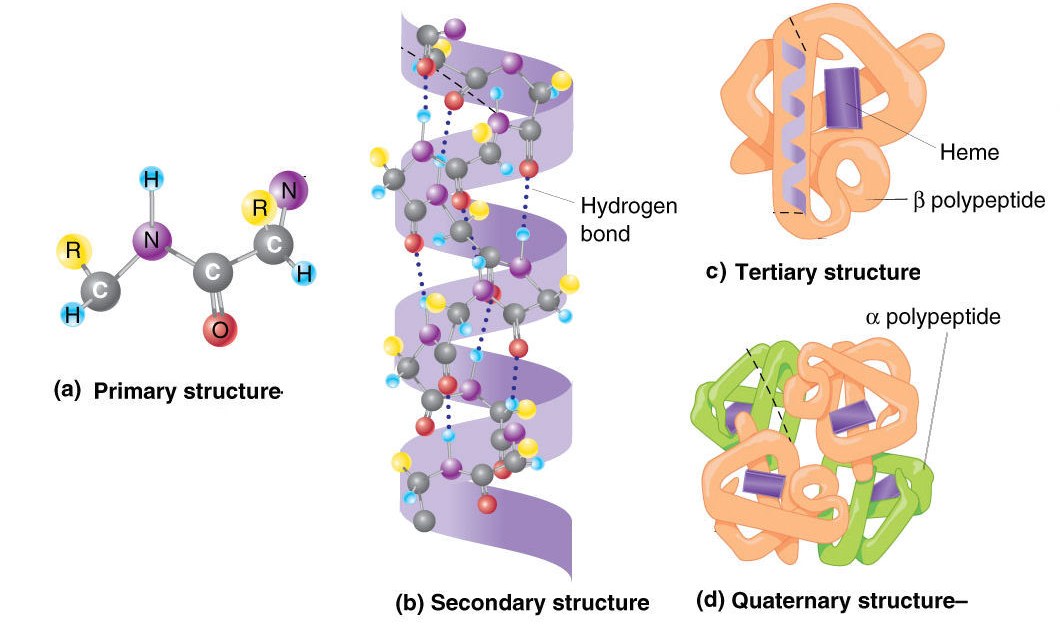
One with a decent graphics card is recommended if possible. You also need a fast enough computer if manipulating and rendering some of these models. Though Blender has a somewhat steep learning curve. It’s assumed you have some basic knowledge of both programs. You can install both freely on any major OS. The basic requirements are the programs PyMOL and Blender. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
